STARCH SYNTHASE5, a Noncanonical Starch Synthase-Like Protein, Promotes Starch Granule Initiation in Arabidopsis
- PMID: 32471861
- PMCID: PMC7401018
- DOI: 10.1105/tpc.19.00946
STARCH SYNTHASE5, a Noncanonical Starch Synthase-Like Protein, Promotes Starch Granule Initiation in Arabidopsis
Abstract
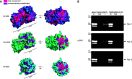
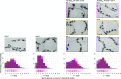
What determines the number of starch granules in plastids is an enigmatic aspect of starch metabolism. Several structurally and functionally diverse proteins have been implicated in the granule initiation process in Arabidopsis (Arabidopsis thaliana), with each protein exerting a varying degree of influence. Here, we show that a conserved starch synthase-like protein, STARCH SYNTHASE5 (SS5), regulates the number of starch granules that form in Arabidopsis chloroplasts. Among the starch synthases, SS5 is most closely related to SS4, a major determinant of granule initiation and morphology. However, unlike SS4 and the other starch synthases, SS5 is a noncanonical isoform that lacks catalytic glycosyltransferase activity. Nevertheless, loss of SS5 reduces starch granule numbers that form per chloroplast in Arabidopsis, and ss5 mutant starch granules are larger than wild-type granules. Like SS4, SS5 has a conserved putative surface binding site for glucans and also interacts with MYOSIN-RESEMBLING CHLOROPLAST PROTEIN, a proposed structural protein influential in starch granule initiation. Phenotypic analysis of a suite of double mutants lacking both SS5 and other proteins implicated in starch granule initiation allows us to propose how SS5 may act in this process.
© 2020 American Society of Plant Biologists. All rights reserved.
Figures








Comment in
-
Which Factors Control Starch Granule Initiation?Plant Cell. 2020 Aug;32(8):2449-2450. doi: 10.1105/tpc.20.00502. Epub 2020 Jun 30. Plant Cell. 2020. PMID: 32605916 Free PMC article. No abstract available.
References
-
- Alonso J.M., et al. (2003). Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301: 653–657. - PubMed
-
- Arrivault S., Guenther M., Ivakov A., Feil R., Vosloh D., van Dongen J.T., Sulpice R., Stitt M.(2009). Use of reverse-phase liquid chromatography, linked to tandem mass spectrometry, to profile the Calvin cycle and other metabolic intermediates in Arabidopsis rosettes at different carbon dioxide concentrations. Plant J. 59: 826–839. - PubMed
-
- Baerenfaller K., Hirsch-Hoffmann M., Svozil J., Hull R., Russenberger D., Bischof S., Lu Q., Gruissem W., Baginsky S.(2011). pep2pro: A new tool for comprehensive proteome data analysis to reveal information about organ-specific proteomes in Arabidopsis thaliana. Integr. Biol. 3: 225–237. - PubMed
-
- Chia T., Chirico M., King R., Ramirez-Gonzalez R., Saccomanno B., Seung D., Simmonds J., Trick M., Uauy C., Verhoeven T., Trafford K.(2020). A carbohydrate-binding protein, B-GRANULE CONTENT 1, influences starch granule size distribution in a dose-dependent manner in polyploid wheat. J. Exp. Bot. 71: 105–115. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

