Release Date: 15 April 2020
- The pygad.GA class accepts a new argument named
fitness_funcwhich accepts a function to be used for calculating the fitness values for the solutions. This allows the project to be customized to any problem by building the right fitness function.
Release Date: 4 May 2020
- The pygad.GA attributes are moved from the class scope to the instance scope.
- Raising an exception for incorrect values of the passed parameters.
- Two new parameters are added to the pygad.GA class constructor
(
init_range_lowandinit_range_high) allowing the user to customize the range from which the genes values in the initial population are selected. - The code object
__code__of the passed fitness function is checked to ensure it has the right number of parameters.
Release Date: 13 May 2020
- The fitness function accepts a new argument named
sol_idxrepresenting the index of the solution within the population. - A new parameter to the pygad.GA class constructor named
initial_populationis supported to allow the user to use a custom initial population to be used by the genetic algorithm. If not None, then the passed population will be used. IfNone, then the genetic algorithm will create the initial population using thesol_per_popandnum_genesparameters. - The parameters
sol_per_popandnum_genesare optional and set toNoneby default. - A new parameter named
callback_generationis introduced in the pygad.GA class constructor. It accepts a function with a single parameter representing the pygad.GA class instance. This function is called after each generation. This helps the user to do post-processing or debugging operations after each generation.
Release Date: 14 May 2020
- The
best_solution()method in the pygad.GA class returns a new output representing the index of the best solution within the population. Now, it returns a total of 3 outputs and their order is: best solution, best solution fitness, and best solution index. Here is an example:
solution, solution_fitness, solution_idx = ga_instance.best_solution()
print("Parameters of the best solution :", solution)
print("Fitness value of the best solution :", solution_fitness, "\n")
print("Index of the best solution :", solution_idx, "\n")- A new attribute named
best_solution_generationis added to the instances of the pygad.GA class. it holds the generation number at which the best solution is reached. It is only assigned the generation number after therun()method completes. Otherwise, its value is -1.Example:
print("Best solution reached after {best_solution_generation} generations.".format(best_solution_generation=ga_instance.best_solution_generation))- The
best_solution_fitnessattribute is renamed tobest_solutions_fitness(plural solution). - Mutation is applied independently for the genes.
Release Date: 17 May 2020
- Adding 2 extra modules (pygad.nn and pygad.gann) for building and training neural networks with the genetic algorithm.
Release Date: 18 May 2020
- The initial value of the
generations_completedattribute of instances from the pygad.GA class is0rather thanNone. - An optional bool parameter named
mutation_by_replacementis added to the constructor of the pygad.GA class. It works only when the selected type of mutation is random (mutation_type="random"). In this case, settingmutation_by_replacement=Truemeans replace the gene by the randomly generated value. IfFalse, then it has no effect and random mutation works by adding the random value to the gene. This parameter should be used when the gene falls within a fixed range and its value must not go out of this range. Here are some examples:
Assume there is a gene with the value 0.5.
If mutation_type="random" and mutation_by_replacement=False,
then the generated random value (e.g. 0.1) will be added to the gene
value. The new gene value is 0.5+0.1=0.6.
If mutation_type="random" and mutation_by_replacement=True, then
the generated random value (e.g. 0.1) will replace the gene value. The
new gene value is 0.1.
Nonevalue could be assigned to themutation_typeandcrossover_typeparameters of the pygad.GA class constructor. WhenNone, this means the step is bypassed and has no action.
Release date: 1 June 2020
- A new module named
pygad.cnnis supported for building convolutional neural networks. - A new module named
pygad.gacnnis supported for training convolutional neural networks using the genetic algorithm. - The
pygad.plot_result()method has 3 optional parameters namedtitle,xlabel, andylabelto customize the plot title, x-axis label, and y-axis label, respectively. - The
pygad.nnmodule supports the softmax activation function. - The name of the
pygad.nn.predict_outputs()function is changed topygad.nn.predict(). - The name of the
pygad.nn.train_network()function is changed topygad.nn.train().
Release date: 5 July 2020
- A new parameter named
delay_after_genis added which accepts a non-negative number specifying the time in seconds to wait after a generation completes and before going to the next generation. It defaults to0.0which means no delay after the generation. - The passed function to the
callback_generationparameter of the pygad.GA class constructor can terminate the execution of the genetic algorithm if it returns the stringstop. This causes therun()method to stop.
One important use case for that feature is to stop the genetic algorithm
when a condition is met before passing though all the generations. The
user may assigned a value of 100 to the num_generations parameter of
the pygad.GA class constructor. Assuming that at generation 50, for
example, a condition is met and the user wants to stop the execution
before waiting the remaining 50 generations. To do that, just make the
function passed to the callback_generation parameter to return the
string stop.
Here is an example of a function to be passed to the
callback_generation parameter which stops the execution if the
fitness value 70 is reached. The value 70 might be the best possible
fitness value. After being reached, then there is no need to pass
through more generations because no further improvement is possible.
def func_generation(ga_instance):
if ga_instance.best_solution()[1] >= 70:
return "stop"Release date: 19 July 2020
- 2 new optional parameters added to the constructor of the
pygad.GAclass which arecrossover_probabilityandmutation_probability.While applying the crossover operation, each parent has a random value generated between 0.0 and 1.0. If this random value is less than or equal to the value assigned to thecrossover_probabilityparameter, then the parent is selected for the crossover operation.For the mutation operation, a random value between 0.0 and 1.0 is generated for each gene in the solution. If this value is less than or equal to the value assigned to themutation_probability, then this gene is selected for mutation. A new optional parameter named
linewidthis added to theplot_result()method to specify the width of the curve in the plot. It defaults to 3.0.Previously, the indices of the genes selected for mutation was randomly generated once for all solutions within the generation. Currently, the genes' indices are randomly generated for each solution in the population. If the population has 4 solutions, the indices are randomly generated 4 times inside the single generation, 1 time for each solution.
Previously, the position of the point(s) for the single-point and two-points crossover was(were) randomly selected once for all solutions within the generation. Currently, the position(s) is(are) randomly selected for each solution in the population. If the population has 4 solutions, the position(s) is(are) randomly generated 4 times inside the single generation, 1 time for each solution.
A new optional parameter named
gene_spaceas added to thepygad.GAclass constructor. It is used to specify the possible values for each gene in case the user wants to restrict the gene values. It is useful if the gene space is restricted to a certain range or to discrete values. For more information, check the More about the ``gene_space` Parameter <https://pygad.readthedocs.io/en/latest/pygad_more.html#more-about-the-gene-space-parameter>`__ section. Thanks to Prof. Tamer A. Farrag for requesting this useful feature.
Release Date: 6 August 2020
- A bug fix in assigning the value to the
initial_populationparameter. - A new parameter named
gene_typeis added to control the gene type. It can be eitherintorfloat. It has an effect only when the parametergene_spaceisNone. - 7 new parameters that accept callback functions:
on_start,on_fitness,on_parents,on_crossover,on_mutation,on_generation, andon_stop.
Release Date: 11 September 2020
- The
learning_rateparameter in thepygad.nn.train()function defaults to 0.01. - Added support of building neural networks for regression using the
new parameter named
problem_type. It is added as a parameter to bothpygad.nn.train()andpygad.nn.predict()functions. The value of this parameter can be either classification or regression to define the problem type. It defaults to classification. - The activation function for a layer can be set to the string
"None"to refer that there is no activation function at this layer. As a result, the supported values for the activation function are"sigmoid","relu","softmax", and"None".
To build a regression network using the pygad.nn module, just do the
following:
- Set the
problem_typeparameter in thepygad.nn.train()andpygad.nn.predict()functions to the string"regression". - Set the activation function for the output layer to the string
"None". This sets no limits on the range of the outputs as it will be from-infinityto+infinity. If you are sure that all outputs will be nonnegative values, then use the ReLU function.
Check the documentation of the pygad.nn module for an example that
builds a neural network for regression. The regression example is also
available at this GitHub
project:
https://github.com/ahmedfgad/NumPyANN
To build and train a regression network using the pygad.gann module,
do the following:
- Set the
problem_typeparameter in thepygad.nn.train()andpygad.nn.predict()functions to the string"regression". - Set the
output_activationparameter in the constructor of thepygad.gann.GANNclass to"None".
Check the documentation of the pygad.gann module for an example that
builds and trains a neural network for regression. The regression
example is also available at this GitHub
project:
https://github.com/ahmedfgad/NeuralGenetic
To build a classification network, either ignore the problem_type
parameter or set it to "classification" (default value). In this
case, the activation function of the last layer can be set to any type
(e.g. softmax).
Release Date: 11 September 2020
- A bug fix when the
problem_typeargument is set toregression.
Release Date: 14 September 2020
- Bug fix to support building and training regression neural networks with multiple outputs.
Release Date: 20 September 2020
- Support of a new module named
kerasgaso that the Keras models can be trained by the genetic algorithm using PyGAD.
Release Date: 3 October 2020
- Bug fix in applying the crossover operation when the
crossover_probabilityparameter is used. Thanks to Eng. Hamada Kassem, Research and Teaching Assistant, Construction Engineering and Management, Faculty of Engineering, Alexandria University, Egypt.
Release Date: 06 December 2020
- The fitness values of the initial population are considered in the
best_solutions_fitnessattribute. - An optional parameter named
save_best_solutionsis added. It defaults toFalse. When it isTrue, then the best solution after each generation is saved into an attribute namedbest_solutions. IfFalse, then no solutions are saved and thebest_solutionsattribute will be empty. - Scattered crossover is supported. To use it, assign the
crossover_typeparameter the value"scattered". - NumPy arrays are now supported by the
gene_spaceparameter. - The following parameters (
gene_type,crossover_probability,mutation_probability,delay_after_gen) can be assigned to a numeric value of any of these data types:int,float,numpy.int,numpy.int8,numpy.int16,numpy.int32,numpy.int64,numpy.float,numpy.float16,numpy.float32, ornumpy.float64.
Release Date: 03 January 2021
- Support of a new module
pygad.torchgato train PyTorch models using PyGAD. Check its documentation. - Support of adaptive mutation where the mutation rate is determined by the fitness value of each solution. Read the Adaptive Mutation section for more details. Also, read this paper: Libelli, S. Marsili, and P. Alba. "Adaptive mutation in genetic algorithms." Soft computing 4.2 (2000): 76-80.
- Before the
run()method completes or exits, the fitness value of the best solution in the current population is appended to thebest_solution_fitnesslist attribute. Note that the fitness value of the best solution in the initial population is already saved at the beginning of the list. So, the fitness value of the best solution is saved before the genetic algorithm starts and after it ends. - When the parameter
parent_selection_typeis set tosss(steady-state selection), then a warning message is printed if the value of thekeep_parentsparameter is set to 0. - More validations to the user input parameters.
- The default value of the
mutation_percent_genesis set to the string"default"rather than the integer 10. This change helps to know whether the user explicitly passed a value to themutation_percent_genesparameter or it is left to its default one. The"default"value is later translated into the integer 10. - The
mutation_percent_genesparameter is no longer accepting the value 0. It must be>0and<=100. - The built-in
warningsmodule is used to show warning messages rather than just using theprint()function. - A new
boolparameter calledsuppress_warningsis added to the constructor of thepygad.GAclass. It allows the user to control whether the warning messages are printed or not. It defaults toFalsewhich means the messages are printed. - A helper method called
adaptive_mutation_population_fitness()is created to calculate the average fitness value used in adaptive mutation to filter the solutions. - The
best_solution()method accepts a new optional parameter calledpop_fitness. It accepts a list of the fitness values of the solutions in the population. IfNone, then thecal_pop_fitness()method is called to calculate the fitness values of the population.
Release Date: 10 January 2021
- In the
gene_spaceparameter, anyNonevalue (regardless of its index or axis), is replaced by a randomly generated number based on the 3 parametersinit_range_low,init_range_high, andgene_type. So, theNonevalue in[..., None, ...]or[..., [..., None, ...], ...]are replaced with random values. This gives more freedom in building the space of values for the genes. - All the numbers passed to the
gene_spaceparameter are casted to the type specified in thegene_typeparameter. - The
numpy.uintdata type is supported for the parameters that accept integer values. - In the
pygad.kerasgamodule, themodel_weights_as_vector()function uses thetrainableattribute of the model's layers to only return the trainable weights in the network. So, only the trainable layers with theirtrainableattribute set toTrue(trainable=True), which is the default value, have their weights evolved. All non-trainable layers with thetrainableattribute set toFalse(trainable=False) will not be evolved. Thanks to Prof. Tamer A. Farrag for pointing about that at GitHub.
Release Date: 15 January 2021
- A bug fix when
save_best_solutions=True. Refer to this issue for more information: ahmedfgad#25
Release Date: 16 February 2021
- In the
gene_spaceargument, the user can use a dictionary to specify the lower and upper limits of the gene. This dictionary must have only 2 items with keyslowandhighto specify the low and high limits of the gene, respectively. This way, PyGAD takes care of not exceeding the value limits of the gene. For a problem with only 2 genes, then usinggene_space=[{'low': 1, 'high': 5}, {'low': 0.2, 'high': 0.81}]means the accepted values in the first gene start from 1 (inclusive) to 5 (exclusive) while the second one has values between 0.2 (inclusive) and 0.85 (exclusive). For more information, please check the Limit the Gene Value Range section of the documentation. - The
plot_result()method returns the figure so that the user can save it. - Bug fixes in copying elements from the gene space.
- For a gene with a set of discrete values (more than 1 value) in the
gene_spaceparameter like[0, 1], it was possible that the gene value may not change after mutation. That is if the current value is 0, then the randomly selected value could also be 0. Now, it is verified that the new value is changed. So, if the current value is 0, then the new value after mutation will not be 0 but 1.
Release Date: 20 February 2021
- 4 new instance attributes are added to hold temporary results after
each generation:
last_generation_fitnessholds the fitness values of the solutions in the last generation,last_generation_parentsholds the parents selected from the last generation,last_generation_offspring_crossoverholds the offspring generated after applying the crossover in the last generation, andlast_generation_offspring_mutationholds the offspring generated after applying the mutation in the last generation. You can access these attributes inside theon_generation()method for example. - A bug fixed when the
initial_populationparameter is used. The bug occurred due to a mismatch between the data type of the array assigned toinitial_populationand the gene type in thegene_typeattribute. Assuming that the array assigned to theinitial_populationparameter is((1, 1), (3, 3), (5, 5), (7, 7))which has typeint. Whengene_typeis set tofloat, then the genes will not be float but casted tointbecause the defined array hasinttype. The bug is fixed by forcing the array assigned toinitial_populationto have the data type in thegene_typeattribute. Check the issue at GitHub: ahmedfgad#27
Thanks to Andrei Rozanski [PhD Bioinformatics Specialist, Department of Tissue Dynamics and Regeneration, Max Planck Institute for Biophysical Chemistry, Germany] for opening my eye to the first change.
Thanks to Marios Giouvanakis, a PhD candidate in Electrical & Computer Engineer, Aristotle University of Thessaloniki (Αριστοτέλειο Πανεπιστήμιο Θεσσαλονίκης), Greece, for emailing me about the second issue.
Release Date: 12 March 2021
- A new
boolparameter calledallow_duplicate_genesis supported. IfTrue, which is the default, then a solution/chromosome may have duplicate gene values. IfFalse, then each gene will have a unique value in its solution. Check the Prevent Duplicates in Gene Values section for more details. - The
last_generation_fitnessis updated at the end of each generation not at the beginning. This keeps the fitness values of the most up-to-date population assigned to thelast_generation_fitnessparameter.
PyGAD 2.14.0 has an issue that is solved in PyGAD 2.14.1. Please consider using 2.14.1 not 2.14.0.
Release Date: 19 May 2021
- Issue
#40
is solved. Now, the
Nonevalue works with thecrossover_typeandmutation_typeparameters: ahmedfgad#40 - The
gene_typeparameter supports accepting alist/tuple/numpy.ndarrayof numeric data types for the genes. This helps to control the data type of each individual gene. Previously, thegene_typecan be assigned only to a single data type that is applied for all genes. For more information, check the More about the ``gene_type` Parameter <https://pygad.readthedocs.io/en/latest/pygad_more.html#more-about-the-gene-type-parameter>`__ section. Thanks to Rainer Engel for asking about this feature in this discussion: ahmedfgad#43 - A new
boolattribute namedgene_type_singleis added to thepygad.GAclass. It isTruewhen there is a single data type assigned to thegene_typeparameter. When thegene_typeparameter is assigned alist/tuple/numpy.ndarray, thengene_type_singleis set toFalse. - The
mutation_by_replacementflag now has no effect ifgene_spaceexists except for the genes withNonevalues. For example, forgene_space=[None, [5, 6]]themutation_by_replacementflag affects only the first gene which hasNonefor its value space. - When an element has a value of
Nonein thegene_spaceparameter (e.g.gene_space=[None, [5, 6]]), then its value will be randomly generated for each solution rather than being generate once for all solutions. Previously, the gene withNonevalue ingene_spaceis the same across all solutions - Some changes in the documentation according to issue #32: ahmedfgad#32
Release Date: 27 May 2021
- Some bug fixes when the
gene_typeparameter is nested. Thanks to Rainer Engel for opening a discussion to report this bug: ahmedfgad#43 (reply in thread)
Rainer Engel helped a lot in suggesting new features and suggesting enhancements in 2.14.0 to 2.14.2 releases.
Release Date: 6 June 2021
- Some bug fixes when setting the
save_best_solutionsparameter toTrue. Previously, the best solution for generationiwas added into thebest_solutionsattribute at generationi+1. Now, thebest_solutionsattribute is updated by each best solution at its exact generation.
Release Date: 17 June 2021
- Control the precision of all genes/individual genes. Thanks to Rainer for asking about this feature: ahmedfgad#43 (comment)
- A new attribute named
last_generation_parents_indicesholds the indices of the selected parents in the last generation. - In adaptive mutation, no need to recalculate the fitness values of
the parents selected in the last generation as these values can be
returned based on the
last_generation_fitnessandlast_generation_parents_indicesattributes. This speeds-up the adaptive mutation. - When a sublist has a value of
Nonein thegene_spaceparameter (e.g.gene_space=[[1, 2, 3], [5, 6, None]]), then its value will be randomly generated for each solution rather than being generated once for all solutions. Previously, a value ofNonein a sublist of thegene_spaceparameter was identical across all solutions. - The dictionary assigned to the
gene_spaceparameter itself or one of its elements has a new key called"step"to specify the step of moving from the start to the end of the range specified by the 2 existing keys"low"and"high". An example is{"low": 0, "high": 30, "step": 2}to have only even values for the gene(s) starting from 0 to 30. For more information, check the More about the ``gene_space` Parameter <https://pygad.readthedocs.io/en/latest/pygad_more.html#more-about-the-gene-space-parameter>`__ section. ahmedfgad#48 - A new function called
predict()is added in both thepygad.kerasgaandpygad.torchgamodules to make predictions. This makes it easier than using custom code each time a prediction is to be made. - A new parameter called
stop_criteriaallows the user to specify one or more stop criteria to stop the evolution based on some conditions. Each criterion is passed asstrwhich has a stop word. The current 2 supported words arereachandsaturate.reachstops therun()method if the fitness value is equal to or greater than a given fitness value. An example forreachis"reach_40"which stops the evolution if the fitness is >= 40.saturatemeans stop the evolution if the fitness saturates for a given number of consecutive generations. An example forsaturateis"saturate_7"which means stop therun()method if the fitness does not change for 7 consecutive generations. Thanks to Rainer for asking about this feature: ahmedfgad#44 - A new bool parameter, defaults to
False, namedsave_solutionsis added to the constructor of thepygad.GAclass. IfTrue, then all solutions in each generation are appended into an attribute calledsolutionswhich is NumPy array. - The
plot_result()method is renamed toplot_fitness(). The users should migrate to the new name as the old name will be removed in the future. - Four new optional parameters are added to the
plot_fitness()function in thepygad.GAclass which arefont_size=14,save_dir=None,color="#3870FF", andplot_type="plot". Usefont_sizeto change the font of the plot title and labels.save_diraccepts the directory to which the figure is saved. It defaults toNonewhich means do not save the figure.colorchanges the color of the plot.plot_typechanges the plot type which can be either"plot"(default),"scatter", or"bar". ahmedfgad#47 - The default value of the
titleparameter in theplot_fitness()method is"PyGAD - Generation vs. Fitness"rather than"PyGAD - Iteration vs. Fitness". - A new method named
plot_new_solution_rate()creates, shows, and returns a figure showing the rate of new/unique solutions explored in each generation. It accepts the same parameters as in theplot_fitness()method. This method only works whensave_solutions=Truein thepygad.GAclass's constructor. - A new method named
plot_genes()creates, shows, and returns a figure to show how each gene changes per each generation. It accepts similar parameters like theplot_fitness()method in addition to thegraph_type,fill_color, andsolutionsparameters. Thegraph_typeparameter can be either"plot"(default),"boxplot", or"histogram".fill_coloraccepts the fill color which works whengraph_typeis either"boxplot"or"histogram".solutionscan be either"all"or"best"to decide whether all solutions or only best solutions are used. - The
gene_typeparameter now supports controlling the precision offloatdata types. For a gene, rather than assigning just the data type likefloat, assign alist/tuple/numpy.ndarraywith 2 elements where the first one is the type and the second one is the precision. For example,[float, 2]forces a gene with a value like0.1234to be0.12. For more information, check the More about the ``gene_type` Parameter <https://pygad.readthedocs.io/en/latest/pygad_more.html#more-about-the-gene-type-parameter>`__ section.
Release Date: 18 June 2021
- Fix a bug when
keep_parentsis set to a positive integer. ahmedfgad#49
Release Date: 18 June 2021
- Fix a bug when using the
kerasgaortorchgamodules. ahmedfgad#51
Release Date: 19 June 2021
- A user-defined function can be passed to the
mutation_type,crossover_type, andparent_selection_typeparameters in thepygad.GAclass to create a custom mutation, crossover, and parent selection operators. Check the User-Defined Crossover, Mutation, and Parent Selection Operators section for more details. ahmedfgad#50
Release Date: 28 September 2021
- The user can use the
tqdmlibrary to show a progress bar. ahmedfgad#50.
import pygad
import numpy
import tqdm
equation_inputs = [4,-2,3.5]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
num_generations = 10000
with tqdm.tqdm(total=num_generations) as pbar:
ga_instance = pygad.GA(num_generations=num_generations,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
on_generation=lambda _: pbar.update(1))
ga_instance.run()
ga_instance.plot_result()But this work does not work if the ga_instance will be pickled (i.e.
the save() method will be called.
ga_instance.save("test")To solve this issue, define a function and pass it to the
on_generation parameter. In the next code, the
on_generation_progress() function is defined which updates the
progress bar.
import pygad
import numpy
import tqdm
equation_inputs = [4,-2,3.5]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution * equation_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
def on_generation_progress(ga):
pbar.update(1)
num_generations = 100
with tqdm.tqdm(total=num_generations) as pbar:
ga_instance = pygad.GA(num_generations=num_generations,
sol_per_pop=5,
num_parents_mating=2,
num_genes=len(equation_inputs),
fitness_func=fitness_func,
on_generation=on_generation_progress)
ga_instance.run()
ga_instance.plot_result()
ga_instance.save("test")- Solved the issue of unequal length between the
solutionsandsolutions_fitnesswhen thesave_solutionsparameter is set toTrue. Now, the fitness of the last population is appended to thesolutions_fitnessarray. ahmedfgad#64 - There was an issue of getting the length of these 4 variables
(
solutions,solutions_fitness,best_solutions, andbest_solutions_fitness) doubled after each call of therun()method. This is solved by resetting these variables at the beginning of therun()method. ahmedfgad#62 - Bug fixes when adaptive mutation is used
(
mutation_type="adaptive"). ahmedfgad#65
Release Date: 2 February 2022
- A new instance attribute called
previous_generation_fitnessadded in thepygad.GAclass. It holds the fitness values of one generation before the fitness values saved in thelast_generation_fitness. - Issue in the
cal_pop_fitness()method in getting the correct indices of the previous parents. This is solved by using the previous generation's fitness saved in the new attributeprevious_generation_fitnessto return the parents' fitness values. Thanks to Tobias Tischhauser (M.Sc. - Mitarbeiter Institut EMS, Departement Technik, OST – Ostschweizer Fachhochschule, Switzerland) for detecting this bug.
Release Date: 2 February 2022
- Validate the fitness value returned from the fitness function. An exception is raised if something is wrong. ahmedfgad#67
Release Date: 8 July 2022
- An issue is solved when the
gene_spaceparameter is given a fixed value. e.g. gene_space=[range(5), 4]. The second gene's value is static (4) which causes an exception. - Fixed the issue where the
allow_duplicate_genesparameter did not work when mutation is disabled (i.e.mutation_type=None). This is by checking for duplicates after crossover directly. ahmedfgad#39 - Solve an issue in the
tournament_selection()method as the indices of the selected parents were incorrect. ahmedfgad#89 - Reuse the fitness values of the previously explored solutions rather
than recalculating them. This feature only works if
save_solutions=True. - Parallel processing is supported. This is by the introduction of a
new parameter named
parallel_processingin the constructor of thepygad.GAclass. Thanks to @windowshopr for opening the issue #78 at GitHub. Check the Parallel Processing in PyGAD section for more information and examples.
Release Date: 9 September 2022
- Raise an exception if the sum of fitness values is zero while either roulette wheel or stochastic universal parent selection is used. ahmedfgad#129
- Initialize the value of the
run_completedproperty toFalse. ahmedfgad#122 - The values of these properties are no longer reset with each call to
the
run()methodself.best_solutions, self.best_solutions_fitness, self.solutions, self.solutions_fitness: ahmedfgad#123. Now, the user can have the flexibility of calling therun()method more than once while extending the data collected after each generation. Another advantage happens when the instance is loaded and therun()method is called, as the old fitness value are shown on the graph alongside with the new fitness values. Read more in this section: Continue without Losing Progress - Thanks Prof. Fernando Jiménez Barrionuevo (Dept. of Information and Communications Engineering, University of Murcia, Murcia, Spain) for editing this comment in the code. https://github.com/ahmedfgad/GeneticAlgorithmPython/commit/5315bbec02777df96ce1ec665c94dece81c440f4
- A bug fixed when
crossover_type=None. - Support of elitism selection through a new parameter named
keep_elitism. It defaults to 1 which means for each generation keep only the best solution in the next generation. If assigned 0, then it has no effect. Read more in this section: Elitism Selection. ahmedfgad#74 - A new instance attribute named
last_generation_elitismadded to hold the elitism in the last generation. - A new parameter called
random_seedadded to accept a seed for the random function generators. Credit to this issue ahmedfgad#70 and Prof. Fernando Jiménez Barrionuevo. Read more in this section: Random Seed. - Editing the
pygad.TorchGAmodule to make sure the tensor data is moved from GPU to CPU. Thanks to Rasmus Johansson for opening this pull request: ahmedfgad/TorchGA#2
Release Date: 19 September 2022
- A big fix when
keep_elitismis used. ahmedfgad#132
Release Date: 14 February 2023
- Remove
numpy.intandnumpy.floatfrom the list of supported data types. ahmedfgad#151 ahmedfgad#152 - Call the
on_crossover()callback function even ifcrossover_typeisNone. ahmedfgad#138 - Call the
on_mutation()callback function even ifmutation_typeisNone. ahmedfgad#138
Release Date: 14 February 2023
- Bug fixes.
Release Date: 22 February 2023
- A new
summary()method is supported to return a Keras-like summary of the PyGAD lifecycle. - A new optional parameter called
fitness_batch_sizeis supported to calculate the fitness in batches. If it is assigned the value1orNone(default), then the normal flow is used where the fitness function is called for each individual solution. If thefitness_batch_sizeparameter is assigned a value satisfying this condition1 < fitness_batch_size <= sol_per_pop, then the solutions are grouped into batches of sizefitness_batch_sizeand the fitness function is called once for each batch. In this case, the fitness function must return a list/tuple/numpy.ndarray with a length equal to the number of solutions passed. ahmedfgad#136. - The
cloudpicklelibrary (https://github.com/cloudpipe/cloudpickle) is used instead of thepicklelibrary to pickle thepygad.GAobjects. This solves the issue of having to redefine the functions (e.g. fitness function). Thecloudpicklelibrary is added as a dependency in therequirements.txtfile. ahmedfgad#159 - Support of assigning methods to these parameters:
fitness_func,crossover_type,mutation_type,parent_selection_type,on_start,on_fitness,on_parents,on_crossover,on_mutation,on_generation, andon_stop. ahmedfgad#92 ahmedfgad#138 - Validating the output of the parent selection, crossover, and mutation functions.
- The built-in parent selection operators return the parent's indices as a NumPy array.
- The outputs of the parent selection, crossover, and mutation operators must be NumPy arrays.
- Fix an issue when
allow_duplicate_genes=True. ahmedfgad#39 - Fix an issue creating scatter plots of the solutions' fitness.
- Sampling from a
set()is no longer supported in Python 3.11. Instead, sampling happens from alist(). ThanksMarco Brennafor pointing to this issue. - The lifecycle is updated to reflect that the new population's fitness is calculated at the end of the lifecycle not at the beginning. ahmedfgad#154 (comment)
- There was an issue when
save_solutions=Truethat causes the fitness function to be called for solutions already explored and have their fitness pre-calculated. ahmedfgad#160 - A new instance attribute named
last_generation_elitism_indicesadded to hold the indices of the selected elitism. This attribute helps to re-use the fitness of the elitism instead of calling the fitness function. - Fewer calls to the
best_solution()method which in turns saves some calls to the fitness function. - Some updates in the documentation to give more details about the
cal_pop_fitness()method. ahmedfgad#79 (comment)
Release Date: 22 February 2023
- Add the cloudpickle library as a dependency.
Release Date 23 February 2023
- Fix an issue when parallel processing was used where the elitism solutions' fitness values are not re-used. ahmedfgad#160 (comment)
Release Date 8 April 2023
- The structure of the library is changed and some methods defined in
the
pygad.pymodule are moved to thepygad.utils,pygad.helper, andpygad.visualizesubmodules. - The
pygad.utils.parent_selectionmodule has a class namedParentSelectionwhere all the parent selection operators exist. Thepygad.GAclass extends this class. - The
pygad.utils.crossovermodule has a class namedCrossoverwhere all the crossover operators exist. Thepygad.GAclass extends this class. - The
pygad.utils.mutationmodule has a class namedMutationwhere all the mutation operators exist. Thepygad.GAclass extends this class. - The
pygad.helper.uniquemodule has a class namedUniquesome helper methods exist to solve duplicate genes and make sure every gene is unique. Thepygad.GAclass extends this class. - The
pygad.visualize.plotmodule has a class namedPlotwhere all the methods that create plots exist. Thepygad.GAclass extends this class. - Support of using the
loggingmodule to log the outputs to both the console and text file instead of using theprint()function. This is by assigning thelogging.Loggerto the newloggerparameter. Check the Logging Outputs for more information. - A new instance attribute called
loggerto save the logger. - The function/method passed to the
fitness_funcparameter accepts a new parameter that refers to the instance of thepygad.GAclass. Check this for an example: Use Functions and Methods to Build Fitness Function and Callbacks. ahmedfgad#163 - Update the documentation to include an example of using functions and methods to calculate the fitness and build callbacks. Check this for more details: Use Functions and Methods to Build Fitness Function and Callbacks. ahmedfgad#92 (comment)
- Validate the value passed to the
initial_populationparameter. - Validate the type and length of the
pop_fitnessparameter of thebest_solution()method. - Some edits in the documentation. ahmedfgad#106
- Fix an issue when building the initial population as (some) genes
have their value taken from the mutation range (defined by the
parameters
random_mutation_min_valandrandom_mutation_max_val) instead of using the parametersinit_range_lowandinit_range_high. - The
summary()method returns the summary as a single-line string. Just log/print the returned string it to see it properly. - The
callback_generationparameter is removed. Use theon_generationparameter instead. - There was an issue when using the
parallel_processingparameter with Keras and PyTorch. As Keras/PyTorch are not thread-safe, thepredict()method gives incorrect and weird results when more than 1 thread is used. ahmedfgad#145 ahmedfgad/TorchGA#5 ahmedfgad/KerasGA#6. Thanks to this StackOverflow answer. - Replace
numpy.floatbyfloatin the 2 parent selection operators roulette wheel and stochastic universal. ahmedfgad#168
Release Date 20 April 2023
- Fix an issue with passing user-defined function/method for parent selection. ahmedfgad#179
Release Date 20 June 2023
- Fix a bug when the initial population has duplciate genes if a nested gene space is used.
- The
gene_spaceparameter can no longer be assigned a tuple. - Fix a bug when the
gene_spaceparameter has a member of typetuple. - A new instance attribute called
gene_space_unpackedwhich has the unpackedgene_space. It is used to solve duplicates. For infinite ranges in thegene_space, they are unpacked to a limited number of values (e.g. 100). - Bug fixes when creating the initial population using
gene_spaceattribute. - When a
dictis used with thegene_spaceattribute, the new gene value was calculated by summing 2 values: 1) the value sampled from thedict2) a random value returned from the random mutation range defined by the 2 parametersrandom_mutation_min_valandrandom_mutation_max_val. This might cause the gene value to exceed the range limit defined in thegene_space. To respect thegene_spacerange, this release only returns the value from thedictwithout summing it to a random value. - Formatting the strings using f-string instead of the
format()method. ahmedfgad#189 - In the
__init__()of thepygad.GAclass, the logged error messages are handled using atry-exceptblock instead of repeating thelogger.error()command. ahmedfgad#189 - A new class named
CustomLoggeris created in thepygad.cnnmodule to create a default logger using theloggingmodule assigned to theloggerattribute. This class is extended in all other classes in the module. The constructors of these classes have a new parameter namedloggerwhich defaults toNone. If no logger is passed, then the default logger in theCustomLoggerclass is used. - Except for the
pygad.nnmodule, theprint()function in all other modules are replaced by theloggingmodule to log messages. - The callback functions/methods
on_fitness(),on_parents(),on_crossover(), andon_mutation()can return values. These returned values override the corresponding properties. The output ofon_fitness()overrides the population fitness. Theon_parents()function/method must return 2 values representing the parents and their indices. The output ofon_crossover()overrides the crossover offspring. The output ofon_mutation()overrides the mutation offspring. - Fix a bug when adaptive mutation is used while
fitness_batch_size>1. ahmedfgad#195 - When
allow_duplicate_genes=Falseand a user-definedgene_spaceis used, it sometimes happen that there is no room to solve the duplicates between the 2 genes by simply replacing the value of one gene by another gene. This release tries to solve such duplicates by looking for a third gene that will help in solving the duplicates. Check this section for more information. - Use probabilities to select parents using the rank parent selection method. ahmedfgad#205
- The 2 parameters
random_mutation_min_valandrandom_mutation_max_valcan accept iterables (list/tuple/numpy.ndarray) with length equal to the number of genes. This enables customizing the mutation range for each individual gene. ahmedfgad#198 - The 2 parameters
init_range_lowandinit_range_highcan accept iterables (list/tuple/numpy.ndarray) with length equal to the number of genes. This enables customizing the initial range for each individual gene when creating the initial population. - The
dataparameter in thepredict()function of thepygad.kerasgamodule can be assigned a data generator. ahmedfgad#115 ahmedfgad#207 - The
predict()function of thepygad.kerasgamodule accepts 3 optional parameters: 1)batch_size=None,verbose=0, andsteps=None. Check documentation of the Keras Model.predict() method for more information. ahmedfgad#207 - The documentation is updated to explain how mutation works when
gene_spaceis used withintorfloatdata types. Check this section. ahmedfgad#198
Release Date 7 September 2023
- A new module
pygad.utils.nsga2is created that has theNSGA2class that includes the functionalities of NSGA-II. The class has these methods: 1)get_non_dominated_set()2)non_dominated_sorting()3)crowding_distance()4)sort_solutions_nsga2(). Check this section for an example. - Support of multi-objective optimization using Non-Dominated Sorting
Genetic Algorithm II (NSGA-II) using the
NSGA2class in thepygad.utils.nsga2module. Just return alist,tuple, ornumpy.ndarrayfrom the fitness function and the library will consider the problem as multi-objective optimization. All the objectives are expected to be maximization. Check this section for an example. - The parent selection methods and adaptive mutation are edited to support multi-objective optimization.
- Two new NSGA-II parent selection methods are supported in the
pygad.utils.parent_selectionmodule: 1) Tournament selection for NSGA-II 2) NSGA-II selection. - The
plot_fitness()method in thepygad.plotmodule has a new optional parameter namedlabelto accept the label of the plots. This is only used for multi-objective problems. Otherwise, it is ignored. It defaults toNoneand accepts alist,tuple, ornumpy.ndarray. The labels are used in a legend inside the plot. - The default color in the methods of the
pygad.plotmodule is changed to the greenish#64f20ccolor. - A new instance attribute named
pareto_frontsadded to thepygad.GAinstances that holds the pareto fronts when solving a multi-objective problem. - The
gene_typeaccepts alist,tuple, ornumpy.ndarrayfor integer data types given that the precision is set toNone(e.g.gene_type=[float, [int, None]]). - In the
cal_pop_fitness()method, the fitness value is re-used ifsave_best_solutions=Trueand the solution is found in thebest_solutionsattribute. These parameters also can help re-using the fitness of a solution instead of calling the fitness function:keep_elitism,keep_parents, andsave_solutions. - The value
99999999999is replaced byfloat('inf')in the 2 methodswheel_cumulative_probs()andstochastic_universal_selection()inside thepygad.utils.parent_selection.ParentSelectionclass. - The
plot_result()method in thepygad.visualize.plot.Plotclass is removed. Instead, please use theplot_fitness()if you did not upgrade yet.
Release Date 29 January 2024
- Solve bugs when multi-objective optimization is used. ahmedfgad#238
- When the
stop_ciiteriaparameter is used with thereachkeyword, then multiple numeric values can be passed when solving a multi-objective problem. For example, if a problem has 3 objective functions, thenstop_criteria="reach_10_20_30"means the GA stops if the fitness of the 3 objectives are at least 10, 20, and 30, respectively. The number values must match the number of objective functions. If a single value found (e.g.stop_criteria=reach_5) when solving a multi-objective problem, then it is used across all the objectives. ahmedfgad#238 - The
delay_after_genparameter is now deprecated and will be removed in a future release. If it is necessary to have a time delay after each generation, then assign a callback function/method to theon_generationparameter to pause the evolution. - Parallel processing now supports calculating the fitness during adaptive mutation. ahmedfgad#201
- The population size can be changed during runtime by changing all the parameters that would affect the size of any thing used by the GA. For more information, check the Change Population Size during Runtime section. ahmedfgad#234
- When a dictionary exists in the
gene_spaceparameter without a step, then mutation occurs by adding a random value to the gene value. The random vaue is generated based on the 2 parametersrandom_mutation_min_valandrandom_mutation_max_val. For more information, check the How Mutation Works with the gene_space Parameter? section. ahmedfgad#229 - Add
objectas a supported data type for int (GA.supported_int_types) and float (GA.supported_float_types). ahmedfgad#174 - Use the
raiseclause instead of thesys.exit(-1)to terminate the execution. ahmedfgad#213 - Fix a bug when multi-objective optimization is used with batch
fitness calculation (e.g.
fitness_batch_sizeset to a non-zero number). - Fix a bug in the
pygad.pyscript when finding the index of the best solution. It does not work properly with multi-objective optimization whereself.best_solutions_fitnesshave multiple columns.
self.best_solution_generation = numpy.where(numpy.array(
self.best_solutions_fitness) == numpy.max(numpy.array(self.best_solutions_fitness)))[0][0]Release Date 17 February 2024
- After the last generation and before the
run()method completes, update the 2 instance attributes: 1)last_generation_parents2)last_generation_parents_indices. This is to keep the list of parents up-to-date with the latest population fitnesslast_generation_fitness. ahmedfgad#275 - 4 methods with names starting with
run_. Their purpose is to keep the main loop inside therun()method clean. Check the Other Methods section for more information.
The PyGAD library is available at PyPI at this page https://pypi.org/project/pygad. PyGAD is built out of a number of open-source GitHub projects. A brief note about these projects is given in the next subsections.
GitHub Link: https://github.com/ahmedfgad/GeneticAlgorithmPython
GeneticAlgorithmPython is the first project which is an open-source Python 3 project for implementing the genetic algorithm based on NumPy.
GitHub Link: https://github.com/ahmedfgad/NumPyANN
NumPyANN builds artificial neural networks in Python 3 using NumPy from scratch. The purpose of this project is to only implement the forward pass of a neural network without using a training algorithm. Currently, it only supports classification and later regression will be also supported. Moreover, only one class is supported per sample.
GitHub Link: https://github.com/ahmedfgad/NeuralGenetic
NeuralGenetic trains neural networks using the genetic algorithm based on the previous 2 projects GeneticAlgorithmPython and NumPyANN.
GitHub Link: https://github.com/ahmedfgad/NumPyCNN
NumPyCNN builds convolutional neural networks using NumPy. The purpose of this project is to only implement the forward pass of a convolutional neural network without using a training algorithm.
GitHub Link: https://github.com/ahmedfgad/CNNGenetic
CNNGenetic trains convolutional neural networks using the genetic algorithm. It uses the GeneticAlgorithmPython project for building the genetic algorithm.
GitHub Link: https://github.com/ahmedfgad/KerasGA
KerasGA trains Keras models using the genetic algorithm. It uses the GeneticAlgorithmPython project for building the genetic algorithm.
GitHub Link: https://github.com/ahmedfgad/TorchGA
TorchGA trains PyTorch models using the genetic algorithm. It uses the GeneticAlgorithmPython project for building the genetic algorithm.
pygad.torchga: https://github.com/ahmedfgad/TorchGA
https://www.linkedin.com/pulse/validation-short-term-parametric-trading-model-genetic-landolfi
https://itchef.ru/articles/397758
https://blog.csdn.net/sinat_38079265/article/details/108449614
If there is an issue using PyGAD, then use any of your preferred option to discuss that issue.
One way is submitting an issue into this GitHub project (github.com/ahmedfgad/GeneticAlgorithmPython) in case something is not working properly or to ask for questions.
If this is not a proper option for you, then check the Contact Us section for more contact details.
PyGAD is actively developed with the goal of building a dynamic library for suporting a wide-range of problems to be optimized using the genetic algorithm.
To ask for a new feature, either submit an issue into this GitHub project (github.com/ahmedfgad/GeneticAlgorithmPython) or send an e-mail to ahmed.f.gad@gmail.com.
Also check the Contact Us section for more contact details.
If you created a project that uses PyGAD, then we can support you by mentioning this project here in PyGAD's documentation.
To do that, please send a message at ahmed.f.gad@gmail.com or check the Contact Us section for more contact details.
Within your message, please send the following details:
- Project title
- Brief description
- Preferably, a link that directs the readers to your project
In this tutorial, we’ll see why mutation with a fixed number of genes is bad, and how to replace it with adaptive mutation. Using the PyGAD Python 3 library, we’ll discuss a few examples that use both random and adaptive mutation.
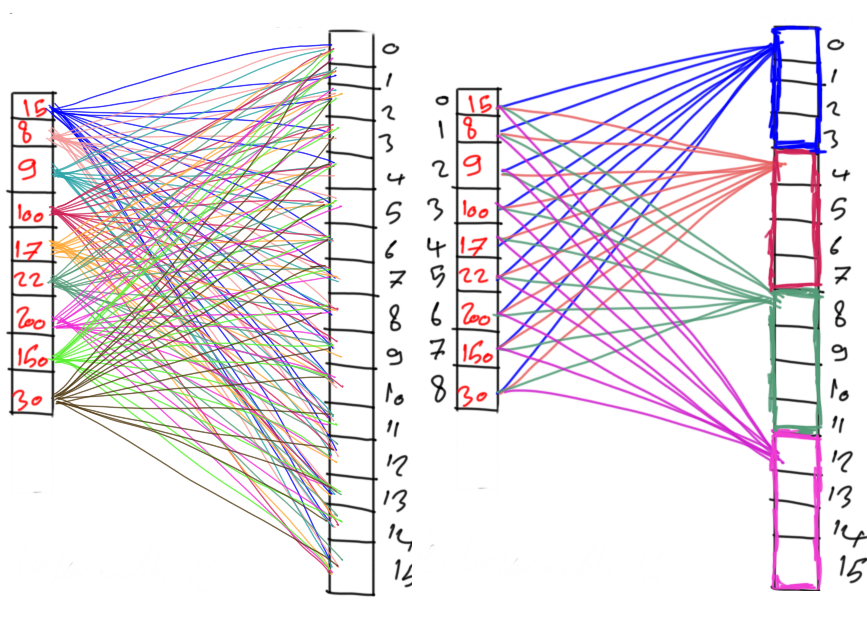
This tutorial discusses how the genetic algorithm is used to cluster data, starting from random clusters and running until the optimal clusters are found. We'll start by briefly revising the K-means clustering algorithm to point out its weak points, which are later solved by the genetic algorithm. The code examples in this tutorial are implemented in Python using the PyGAD library.
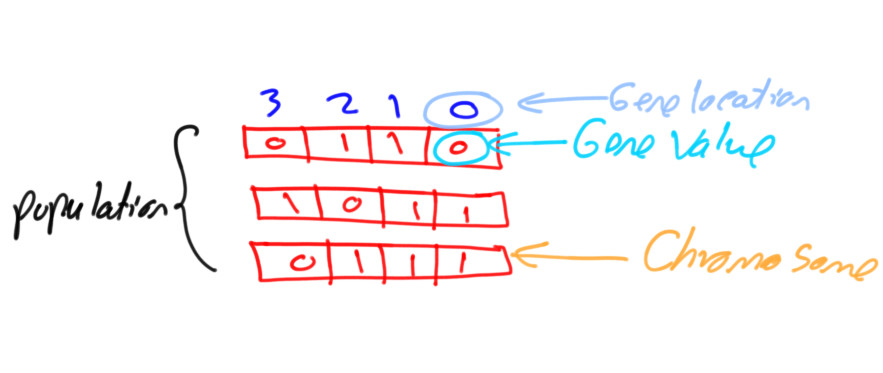
Depending on the nature of the problem being optimized, the genetic algorithm (GA) supports two different gene representations: binary, and decimal. The binary GA has only two values for its genes, which are 0 and 1. This is easier to manage as its gene values are limited compared to the decimal GA, for which we can use different formats like float or integer, and limited or unlimited ranges.
This tutorial discusses how the PyGAD library supports the two GA representations, binary and decimal.
This tutorial introduces PyGAD, an open-source Python library for implementing the genetic algorithm and training machine learning algorithms. PyGAD supports 19 parameters for customizing the genetic algorithm for various applications.
Within this tutorial we'll discuss 5 different applications of the genetic algorithm and build them using PyGAD.
The genetic algorithm (GA) is a biologically-inspired optimization algorithm. It has in recent years gained importance, as it’s simple while also solving complex problems like travel route optimization, training machine learning algorithms, working with single and multi-objective problems, game playing, and more.
Deep neural networks are inspired by the idea of how the biological brain works. It’s a universal function approximator, which is capable of simulating any function, and is now used to solve the most complex problems in machine learning. What’s more, they’re able to work with all types of data (images, audio, video, and text).
Both genetic algorithms (GAs) and neural networks (NNs) are similar, as both are biologically-inspired techniques. This similarity motivates us to create a hybrid of both to see whether a GA can train NNs with high accuracy.
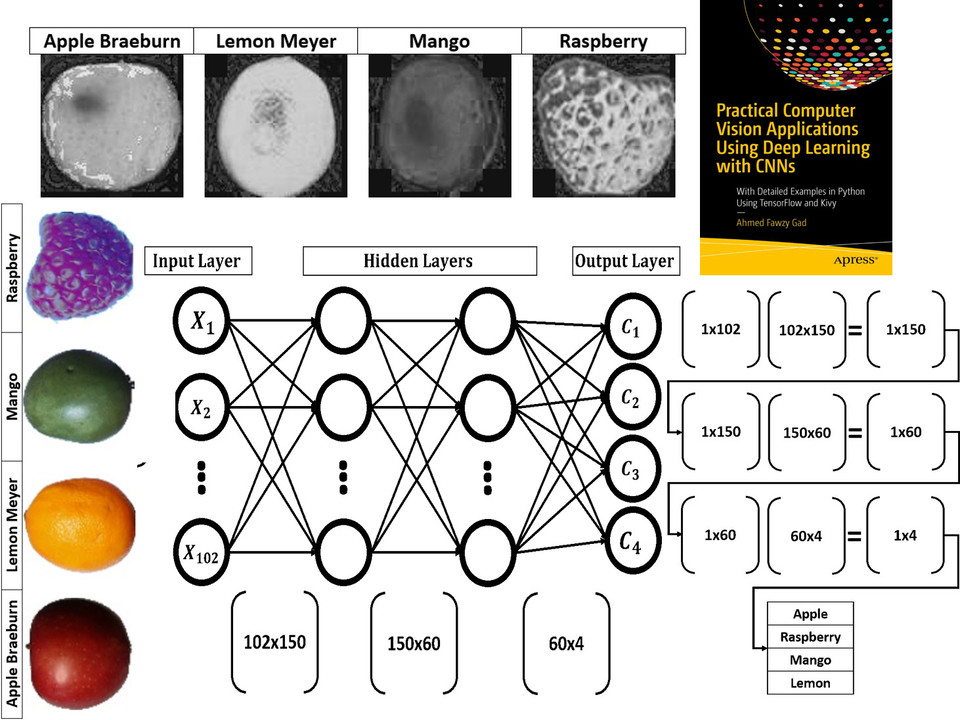
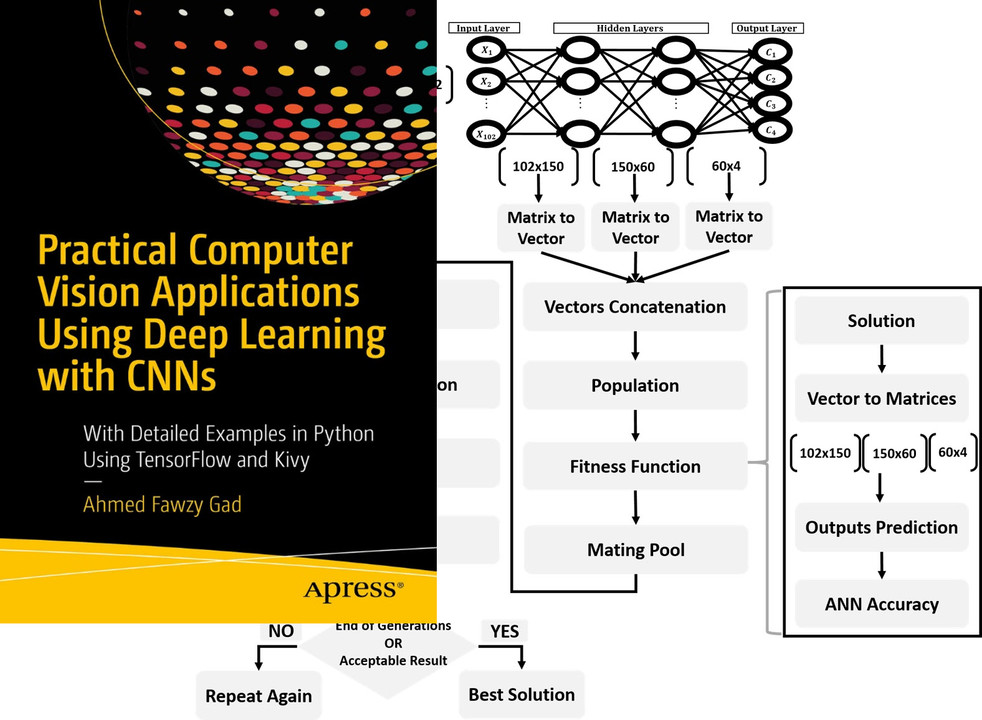
This tutorial uses PyGAD, a Python library that supports building and training NNs using a GA. PyGAD offers both classification and regression NNs.
In this tutorial we'll see how to build a game-playing agent using only the genetic algorithm to play a game called CoinTex, which is developed in the Kivy Python framework. The objective of CoinTex is to collect the randomly distributed coins while avoiding collision with fire and monsters (that move randomly). The source code of CoinTex can be found on GitHub.
The genetic algorithm is the only AI used here; there is no other machine/deep learning model used with it. We'll implement the genetic algorithm using PyGad. This tutorial starts with a quick overview of CoinTex followed by a brief explanation of the genetic algorithm, and how it can be used to create the playing agent. Finally, we'll see how to implement these ideas in Python.
The source code of the genetic algorithm agent is available here, and you can download the code used in this tutorial from here.
PyGAD is an open-source Python library for building the genetic algorithm and training machine learning algorithms. It offers a wide range of parameters to customize the genetic algorithm to work with different types of problems.
PyGAD has its own modules that support building and training neural networks (NNs) and convolutional neural networks (CNNs). Despite these modules working well, they are implemented in Python without any additional optimization measures. This leads to comparatively high computational times for even simple problems.
The latest PyGAD version, 2.8.0 (released on 20 September 2020), supports a new module to train Keras models. Even though Keras is built in Python, it's fast. The reason is that Keras uses TensorFlow as a backend, and TensorFlow is highly optimized.
This tutorial discusses how to train Keras models using PyGAD. The discussion includes building Keras models using either the Sequential Model or the Functional API, building an initial population of Keras model parameters, creating an appropriate fitness function, and more.
PyGAD is a genetic algorithm Python 3 library for solving optimization problems. One of these problems is training machine learning algorithms.
PyGAD has a module called pygad.kerasga. It trains Keras models using the genetic algorithm. On January 3rd, 2021, a new release of PyGAD 2.10.0 brought a new module called pygad.torchga to train PyTorch models. It’s very easy to use, but there are a few tricky steps.
So, in this tutorial, we’ll explore how to use PyGAD to train PyTorch models.
Cómo los algoritmos genéticos pueden competir con el descenso de gradiente y el backprop
Bien que la manière standard d'entraîner les réseaux de neurones soit la descente de gradient et la rétropropagation, il y a d'autres joueurs dans le jeu. L'un d'eux est les algorithmes évolutionnaires, tels que les algorithmes génétiques.
Utiliser un algorithme génétique pour former un réseau de neurones simple pour résoudre le OpenAI CartPole Jeu. Dans cet article, nous allons former un simple réseau de neurones pour résoudre le OpenAI CartPole . J'utiliserai PyTorch et PyGAD .
Cómo los algoritmos genéticos pueden competir con el descenso de gradiente y el backprop
Aunque la forma estandar de entrenar redes neuronales es el descenso de gradiente y la retropropagacion, hay otros jugadores en el juego, uno de ellos son los algoritmos evolutivos, como los algoritmos geneticos.
Usa un algoritmo genetico para entrenar una red neuronal simple para resolver el Juego OpenAI CartPole. En este articulo, entrenaremos una red neuronal simple para resolver el OpenAI CartPole . Usare PyTorch y PyGAD .
파이썬에서 genetic algorithm을 사용하는 패키지들을 다 사용해보진 않았지만, 확장성이 있어보이고, 시도할 일이 있어서 살펴봤다.
이 패키지에서 가장 인상 깊었던 것은 neural network에서 hyper parameter 탐색을 gradient descent 방식이 아닌 GA로도 할 수 있다는 것이다.
개인적으로 이 부분이 어느정도 초기치를 잘 잡아줄 수 있는 역할로도 쓸 수 있고, Loss가 gradient descent 하기 어려운 구조에서 대안으로 쓸 수 있을 것으로도 생각된다.
일단 큰 흐름은 다음과 같이 된다.
사실 완전히 흐름이나 각 parameter에 대한 이해는 부족한 상황
This is a translation of an original English tutorial published at Paperspace: How To Train Keras Models Using the Genetic Algorithm with PyGAD
PyGAD, genetik algoritma oluşturmak ve makine öğrenimi algoritmalarını eğitmek için kullanılan açık kaynaklı bir Python kitaplığıdır. Genetik algoritmayı farklı problem türleri ile çalışacak şekilde özelleştirmek için çok çeşitli parametreler sunar.
PyGAD, sinir ağları (NN’ler) ve evrişimli sinir ağları (CNN’ler) oluşturmayı ve eğitmeyi destekleyen kendi modüllerine sahiptir. Bu modüllerin iyi çalışmasına rağmen, herhangi bir ek optimizasyon önlemi olmaksızın Python’da uygulanırlar. Bu, basit problemler için bile nispeten yüksek hesaplama sürelerine yol açar.
En son PyGAD sürümü 2.8.0 (20 Eylül 2020'de piyasaya sürüldü), Keras modellerini eğitmek için yeni bir modülü destekliyor. Keras Python’da oluşturulmuş olsa da hızlıdır. Bunun nedeni, Keras’ın arka uç olarak TensorFlow kullanması ve TensorFlow’un oldukça optimize edilmiş olmasıdır.
Bu öğreticide, PyGAD kullanılarak Keras modellerinin nasıl eğitileceği anlatılmaktadır. Tartışma, Sıralı Modeli veya İşlevsel API’yi kullanarak Keras modellerini oluşturmayı, Keras model parametrelerinin ilk popülasyonunu oluşturmayı, uygun bir uygunluk işlevi oluşturmayı ve daha fazlasını içerir.
Tensorflow alapozó 10. Neurális hálózatok tenyésztése genetikus algoritmussal PyGAD és OpenAI Gym használatával
Hogy kontextusba helyezzem a genetikus algoritmusokat, ismételjük kicsit át, hogy hogyan működik a gradient descent és a backpropagation, ami a neurális hálók tanításának általános módszere. Az erről írt cikkemet itt tudjátok elolvasni.
A hálózatok tenyésztéséhez a PyGAD nevű programkönyvtárat használjuk, így mindenek előtt ezt kell telepítenünk, valamint a Tensorflow-t és a Gym-et, amit Colabban már eleve telepítve kapunk.
Maga a PyGAD egy teljesen általános genetikus algoritmusok futtatására képes rendszer. Ennek a kiterjesztése a KerasGA, ami az általános motor Tensorflow (Keras) neurális hálókon történő futtatását segíti. A 47. sorban létrehozott KerasGA objektum ennek a kiterjesztésnek a része és arra szolgál, hogy a paraméterként átadott modellből a második paraméterben megadott számosságú populációt hozzon létre. Mivel a hálózatunk 386 állítható paraméterrel rendelkezik, ezért a DNS-ünk itt 386 elemből fog állni. A populáció mérete 10 egyed, így a kezdő populációnk egy 10x386 elemű mátrix lesz. Ezt adjuk át az 51. sorban az initial_population paraméterben.
PyGAD — это библиотека для имплементации генетического алгоритма. Кроме того, библиотека предоставляет доступ к оптимизированным реализациям алгоритмов машинного обучения. PyGAD разрабатывали на Python 3.
Библиотека PyGAD поддерживает разные типы скрещивания, мутации и селекции родителя. PyGAD позволяет оптимизировать проблемы с помощью генетического алгоритма через кастомизацию целевой функции.
Кроме генетического алгоритма, библиотека содержит оптимизированные имплементации алгоритмов машинного обучения. На текущий момент PyGAD поддерживает создание и обучение нейросетей для задач классификации.
Библиотека находится в стадии активной разработки. Создатели планируют добавление функционала для решения бинарных задач и имплементации новых алгоритмов.
PyGAD разрабатывали на Python 3.7.3. Зависимости включают в себя NumPy для создания и манипуляции массивами и Matplotlib для визуализации. Один из изкейсов использования инструмента — оптимизация весов, которые удовлетворяют заданной функции.
A number of research papers used PyGAD and here are some of them:
- Alberto Meola, Manuel Winkler, Sören Weinrich, Metaheuristic optimization of data preparation and machine learning hyperparameters for prediction of dynamic methane production, Bioresource Technology, Volume 372, 2023, 128604, ISSN 0960-8524.
- Jaros, Marta, and Jiri Jaros. "Performance-Cost Optimization of Moldable Scientific Workflows."
- Thorat, Divya. "Enhanced genetic algorithm to reduce makespan of multiple jobs in map-reduce application on serverless platform". Diss. Dublin, National College of Ireland, 2020.
- Koch, Chris, and Edgar Dobriban. "AttenGen: Generating Live Attenuated Vaccine Candidates using Machine Learning." (2021).
- Bhardwaj, Bhavya, et al. "Windfarm optimization using Nelder-Mead and Particle Swarm optimization." 2021 7th International Conference on Electrical Energy Systems (ICEES). IEEE, 2021.
- Bernardo, Reginald Christian S. and J. Said. “Towards a model-independent reconstruction approach for late-time Hubble data.” (2021).
- Duong, Tri Dung, Qian Li, and Guandong Xu. "Prototype-based Counterfactual Explanation for Causal Classification." arXiv preprint arXiv:2105.00703 (2021).
- Farrag, Tamer Ahmed, and Ehab E. Elattar. "Optimized Deep Stacked Long Short-Term Memory Network for Long-Term Load Forecasting." IEEE Access 9 (2021): 68511-68522.
- Antunes, E. D. O., Caetano, M. F., Marotta, M. A., Araujo, A., Bondan, L., Meneguette, R. I., & Rocha Filho, G. P. (2021, August). Soluções Otimizadas para o Problema de Localização de Máxima Cobertura em Redes Militarizadas 4G/LTE. In Anais do XXVI Workshop de Gerência e Operação de Redes e Serviços (pp. 152-165). SBC.
- M. Yani, F. Ardilla, A. A. Saputra and N. Kubota, "Gradient-Free Deep Q-Networks Reinforcement learning: Benchmark and Evaluation," 2021 IEEE Symposium Series on Computational Intelligence (SSCI), 2021, pp. 1-5, doi: 10.1109/SSCI50451.2021.9659941.
- Yani, Mohamad, and Naoyuki Kubota. "Deep Convolutional Networks with Genetic Algorithm for Reinforcement Learning Problem."
- Mahendra, Muhammad Ihza, and Isman Kurniawan. "Optimizing Convolutional Neural Network by Using Genetic Algorithm for COVID-19 Detection in Chest X-Ray Image." 2021 International Conference on Data Science and Its Applications (ICoDSA). IEEE, 2021.
- Glibota, Vjeko. Umjeravanje mikroskopskog prometnog modela primjenom genetskog algoritma. Diss. University of Zagreb. Faculty of Transport and Traffic Sciences. Division of Intelligent Transport Systems and Logistics. Department of Intelligent Transport Systems, 2021.
- Zhu, Mingda. Genetic Algorithm-based Parameter Identification for Ship Manoeuvring Model under Wind Disturbance. MS thesis. NTNU, 2021.
- Abdalrahman, Ahmed, and Weihua Zhuang. "Dynamic pricing for differentiated pev charging services using deep reinforcement learning." IEEE Transactions on Intelligent Transportation Systems (2020).
https://rodriguezanton.com/identifying-contact-states-for-2d-objects-using-pygad-and/
https://torvaney.github.io/projects/t9-optimised
There are different resources that can be used to get started with the genetic algorithm and building it in Python.
To start with coding the genetic algorithm, you can check the tutorial titled Genetic Algorithm Implementation in Python available at these links:
This tutorial is prepared based on a previous version of the project but it still a good resource to start with coding the genetic algorithm.
Get started with the genetic algorithm by reading the tutorial titled Introduction to Optimization with Genetic Algorithm which is available at these links:
Read about building neural networks in Python through the tutorial titled Artificial Neural Network Implementation using NumPy and Classification of the Fruits360 Image Dataset available at these links:
Read about training neural networks using the genetic algorithm through the tutorial titled Artificial Neural Networks Optimization using Genetic Algorithm with Python available at these links:
To start with coding the genetic algorithm, you can check the tutorial titled Building Convolutional Neural Network using NumPy from Scratch available at these links:
This tutorial) is prepared based on a previous version of the project but it still a good resource to start with coding CNNs.
Get started with the genetic algorithm by reading the tutorial titled Derivation of Convolutional Neural Network from Fully Connected Network Step-By-Step which is available at these links:
You can also check my book cited as Ahmed Fawzy Gad 'Practical Computer Vision Applications Using Deep Learning with CNNs'. Dec. 2018, Apress, 978-1-4842-4167-7 which discusses neural networks, convolutional neural networks, deep learning, genetic algorithm, and more.
Find the book at these links:
- E-mail: ahmed.f.gad@gmail.com
- Amazon Author Page
- Heartbeat
- Paperspace
- KDnuggets
- TowardsDataScience
- GitHub
Thank you for using PyGAD :)